AI, Nutrition, and Healthspan: why clinicians can't ignore this new triad
Artificial intelligence has transitioned from a promising concept to a transformative force in healthcare, fundamentally altering how we measure, predict, and manage aging, nutrition, and chronic disease risk across the lifespan. Deep learning and generative AI models now power sophisticated "aging clocks" that estimate biological age from diverse data sources including blood biomarkers, epigenetic markers, and multi-omics datasets, offering a dynamic assessment of healthspan rather than simply tallying chronological years. These computational tools can identify patterns linked to frailty and multimorbidity years before clinical symptoms manifest, creating unprecedented opportunities for earlier, nutrition-focused prevention strategies. This article provides a comprehensive review of AI applications at the intersection of nutrition science, aging research, and clinical practice, synthesizing evidence from recent systematic reviews, randomized controlled trials, and regulatory frameworks to guide evidence-based implementation.
Alessandro Drago
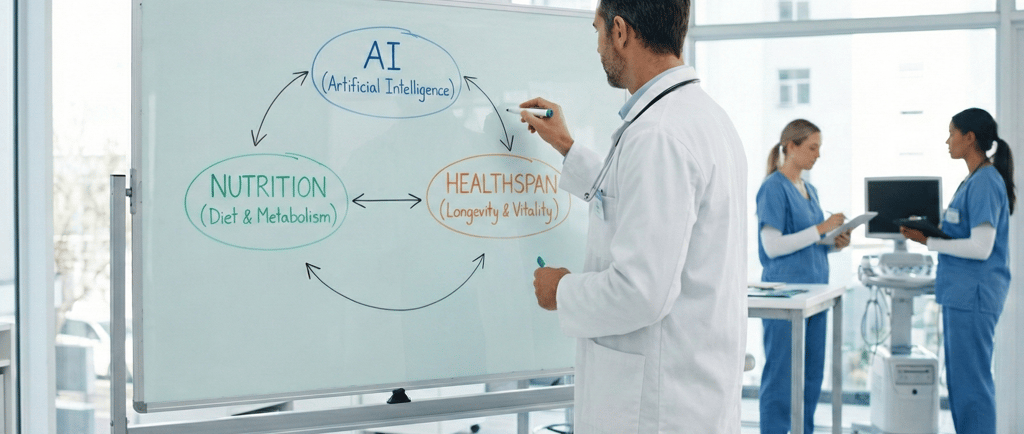
Deep Learning Architectures in Aging Research
Evolution of Biological Age Estimation
The application of deep learning to aging research began in the 2015–2016 period with the development of the first Deep Aging Clocks (DACs) by Zhavoronkov and colleagues. These pioneering models utilized feed-forward deep neural networks trained on blood biochemistry data from reasonably healthy individuals, achieving a mean absolute error of approximately 5.5 years in estimating chronological age. This foundational work established that machine learning algorithms are capable of capturing complex, non-linear relationships among multiple biomarkers that reflect biological aging processes more accurately than chronological age alone.
Modern aging clocks have evolved substantially in both methodology and accuracy. Epigenetic clocks, which measure DNA methylation patterns at specific CpG sites, have emerged as particularly powerful tools. The Horvath clock, one of the most widely used first-generation epigenetic clocks, demonstrates accuracy across multiple tissue types. The Hannum clock, specifically optimized for blood samples, shows particular utility in blood-based health studies and highlights strong associations with clinical markers such as BMI, cardiovascular health, immune function, and chronic conditions. Second-generation clocks, such as PhenoAge, integrate DNA methylation data with nine key clinical biomarkers (including albumin, creatinine, glucose, C-reactive protein, lymphocyte percentage, mean cell volume, red cell distribution width, alkaline phosphatase, and white blood cell count), increasing sensitivity to individual variations and improving the accuracy of biological age estimation.
Recent validation studies have examined 14 alternative aging indicators, including eight epigenetic biomarkers, BioAge composites from clinical laboratory parameters, brain age, skin age, subjective age, and future health horizon. These studies demonstrate moderate associations within categories, supporting the multidimensional nature of biological aging. A comprehensive platform called TranslAGE now harmonizes 179 human blood DNA methylation datasets with pre-calculated scores for 41 epigenetic biomarker models across more than 42,000 samples, enabling systematic validation through a STAR framework (Stability, Treatment response, Associations, Risk). This standardization is critical for advancing the clinical translation of aging biomarkers.
The ENABL age framework
The ENABL Age (ExplaiNAble BioLogical Age) framework represents a significant advancement in addressing the trade-off between accuracy and interpretability that has historically limited the clinical adoption of aging clocks. Developed using datasets from the UK Biobank and the National Health and Nutrition Examination Survey (NHANES), ENABL Age combines machine learning models with explainable artificial intelligence (XAI) methods to both accurately estimate biological age and provide individualized explanations regarding contributing risk factors.
The ENABL Age clock demonstrates a strong correlation with chronological age (r=0.7867, p<0.0001 for UK Biobank; r=0.7126, p<0.0001 for NHANES) while achieving superior mortality prediction compared to existing clocks. Groups identified as "unhealthy" by ENABL Age—meaning their biological age exceeds their chronological age—showed log hazard ratios approximately 3 to 12 times higher than healthy groups. The model achieves high predictive power for mortality, with AUROC values of 0.8179 for 5-year mortality and 0.8115 for 10-year mortality on the UK Biobank dataset, and 0.8935 and 0.9107 respectively on NHANES.
Crucially, ENABL Age provides two accessible versions: ENABL Age-L, based on blood tests, and ENABL Age-Q, based on questionnaire characteristics. Individualized explanations decompose scores into contributing risk factors, revealing that modifiable factors like body composition, blood glucose control, smoking status, physical activity levels, and diet quality can be genuinely prioritized in personalized counseling rather than treated as a universal checklist. For clinicians and dietitians, this translates into actionable intelligence that enables a shift from reactive sick care to proactive healthspan management, provided that human judgment remains firmly in control of clinical decision-making.
AI-Enhanced Dietary Assessment Technologies
Limitations of Traditional Methods
Traditional dietary assessment methods (including 24-hour recalls, food diaries, and food frequency questionnaires) have long been recognized as vulnerable to systematic biases. These methods suffer from memory lapses, portion estimation errors, social desirability bias, and underreporting; phenomena particularly pronounced in older populations. Research demonstrates that these limitations compromise the reliability and validity of dietary data, consequently undermining the robustness of associations between dietary patterns and health outcomes.
AI-Powered Image Recognition and Wearable Technologies
Recent systematic reviews demonstrate that AI-based image recognition, smartphone applications, and wearable sensors can substantially improve portion estimates, automate nutrient calculations, and reduce user burden. A comprehensive scoping review of AI applications for measuring food and nutrient intake identified 25 studies published between 2010 and 2023, examining various input data types including food images, sound and jaw motion data from wearable devices, and text data. Deep learning approaches, particularly convolutional neural networks (CNNs), have achieved food detection accuracies ranging from 74% to 99.85%.
Nutrient estimation, however, presents greater challenges. RGB-D (Red, Green, Blue-Depth) fusion networks have achieved mean absolute errors of approximately 15% in calorie estimation, while sound-based classification models have demonstrated up to 94% accuracy in detecting food intake based on jaw motion and chewing patterns. A systematic review comparing AI-based digital image dietary assessment methods with human estimation found that relative errors for volume and calorie estimates suggest AI methods align with human accuracy, with the potential to exceed it. However, images of simpler foods (such as a single apple) consistently showed lower relative error rates compared to complex mixed dishes.
Evaluations of popular nutrition apps reveal substantial variability in quality and accuracy. Among 18 evaluated apps, Noom scored highest on both the Mobile App Rating Scale (MARS, mean=4.44) and the ABACUS for behavior change potential (21/21). Among manual food-logging apps tested with standardized diets, energy estimates showed systematic bias, with overestimations for Western diets and underestimations for Asian diets. Among AI-enabled food image recognition apps, MyFitnessPal and Fastic demonstrated the highest accuracy, though automatic energy estimations remained substantially inaccurate. These findings underscore that while AI offers significant advantages in improving measurement accuracy and enabling real-time monitoring, challenges remain in adapting to diverse food types, ensuring algorithmic fairness across different cuisines, and addressing data privacy concerns.
Continuous Glucose Monitoring and AI Integration
The integration of continuous glucose monitoring (CGM) with AI-based analytics represents a paradigm shift in personalized nutrition, particularly for the management of diabetes and obesity. CGM devices provide unprecedented continuous, real-time data on glucose dynamics, and when combined with AI-driven analytical tools, enable truly personalized dietary interventions.
Machine learning models incorporating CGM data, meal content, lifestyle factors, biochemical parameters, and microbiome data have demonstrated superior predictive accuracy for postprandial glycemic responses. Studies show that AI can analyze CGM data to identify specific foods causing blood glucose fluctuations and provide optimized dietary recommendations with a precision previously unattainable. Deep neural networks combined with explainable AI methods can analyze multiple factors, including pre-meal blood glucose and insulin dosage, to accurately predict postprandial glucose levels. Notably, AI-guided meal planning based on CGM data achieved a 27% reduction in postprandial glucose spikes in type 2 diabetes patients, allowing healthcare teams to implement timely interventions with personalized feedback, including glucose anomaly alerts.
Clinical Efficacy of AI-Generated Dietary Interventions
Evidence from Randomized Controlled Trials
Evidence from systematic reviews demonstrates that AI-generated meal plans are beginning to outperform conventional dietary advice in specific clinical conditions, particularly diabetes and irritable bowel syndrome (IBS). A systematic review following PRISMA guidelines identified 11 studies (five randomized controlled trials, five pre-post designs, and one cross-sectional analysis) published between 2015 and 2024. Most AI methods employed were based on machine learning, including conventional algorithms, deep learning, and hybrid approaches integrated with IoT systems. These interventions led to improved glycemic control, strengthened metabolic health, and positive psychological outcomes compared to standard diets, with evidence quality rated from moderate to high.
Systems integrating microbiome profiles, metabolic markers, and lifestyle inputs have produced clinically meaningful improvements in blood glucose control and IBS symptoms compared to conventional guideline-based diets. This demonstrates how personalization can enrich, rather than replace, the classical principles of nutrition science.
Applications in Gastroenterology and Hepatology
A recent narrative review of the literature examined AI-guided personalized nutrition in gastroenterology and hepatology, highlighting its transformative potential while acknowledging implementation challenges. For inflammatory bowel disease and IBS, AI allows for microbiome-guided diet personalization by capturing the dynamic interplay between microbial ecosystems, dietary exposures, and host physiology. Compared to conventional dietary guidance, AI-generated plans led to significantly improved glycemic control and favorable changes in gut microbial composition for patients with metabolic dysfunction-associated steatotic liver disease.
For professionals, these platforms function optimally as decision-support tools that suggest tailored options. Clinicians retain ultimate responsibility for clinical appropriateness, facilitating behavioral change, and follow-up care, thus preserving the essential human elements of nutritional counseling such as empathy, motivation, and cultural sensitivity.
AI in Hospital Nutrition Workflows and Malnutrition Prediction
Predictive Models for Malnutrition Risk
AI is increasingly integrated into hospital nutrition workflows to identify patients at risk of malnutrition, muscle loss, or complications before these problems become clinically apparent. Machine learning models trained on electronic health records utilize variables such as BMI, recent weight loss, inflammatory markers, comorbidities, functional scores, albumin levels, and even psycho-social factors like loneliness. These models can predict adverse outcomes more accurately than traditional screening tools alone.
A systematic literature review, however, noted that over 90% of AI models remain unused in daily clinical practice despite demonstrated efficacy. Supervised learning models are the most widespread approach, with disease-related malnutrition as the primary category. The review highlighted critical gaps between research and clinical implementation due to a lack of standardized protocols, ethical concerns regarding bias and privacy, limited validation in diverse populations, and insufficient integration into health information systems.
Specialized Prediction Models for Vulnerable Populations
Several specialized AI models have demonstrated particular utility for vulnerable populations. An interpretable machine learning model for predicting malnutrition in post-stroke patients was developed and validated on multicenter data. Among eight tested models, the CatBoost algorithm achieved superior performance with high predictive accuracy.
For sarcopenia prediction, a personalized model for older European adults with arthritis was developed using longitudinal data spanning 12 years. The model captures long-term sarcopenia progression trajectories, supporting dynamic risk assessment. The integration of interpretability analyses based on SHAP provides transparent insights, allowing clinicians to understand which specific factors drive individual risk predictions. Furthermore, studies on hospitalized elderly patients have demonstrated that machine learning-based approaches can improve the prediction of adverse outcomes including infections, pressure ulcers, falls, and mortality. These models enable earlier dietary assessment, more strategic use of supplements, and better allocation of clinical resources.
Regulatory Landscape and Ethical Frameworks
The European Union AI Act
The rapid proliferation of AI in healthspan and nutrition amplifies the need for critical thinking and regulatory awareness. The European Union AI Act (Regulation EU 2024/1689) entered into force on August 1, 2024, establishing a comprehensive, risk-based legal framework. This legislation treats most AI-enabled medical devices and clinical decision-support systems as "high-risk," requiring strict standards for data quality, transparency, human oversight, and continuous monitoring.
Under the AI Act, any AI system intended as a safety component of a product subject to medical device regulations qualifies automatically as high-risk. This classification includes virtually all AI-enabled medical software, computer-aided diagnosis algorithms, and triage tools. The regulation imposes obligations such as continuous risk management, bias control, and incident reporting within 15 days. Human oversight requirements mandate providers to create mechanisms allowing clinicians to override or contest AI outputs, while deployers must ensure AI literacy and performance monitoring. Penalties for non-compliance can reach €35 million or 7% of global annual turnover, placing AI compliance on the same strategic level as GDPR.
The EFSA Roadmap on AI for Food Safety
In parallel, the European Food Safety Authority (EFSA) launched its roadmap for AI in risk assessment. The project aims to increase the accessibility of scientific evidence and improve the reliability of processes by applying human-centric AI in coexistence with human expertise by 2027.
EFSA's approach strategically focuses on high-volume, low-judgment tasks, such as terminology normalization and literature screening, maximizing impact while reducing risk. The emphasis on validation against gold standards and expert oversight remains central, underscoring that AI serves as an augmentation rather than complete automation.
Implications for Nutrition Professionals
For nutrition professionals, understanding the origin, validation, and limitations of AI tools is becoming as fundamental as traditional clinical skills. Practitioners must develop competencies in evaluating data provenance and quality, validation methodology, the explainability of recommendations, and integration into clinical workflows. Professional organizations must therefore develop educational programs that equip nutritionists with the necessary AI literacy, enabling them to interpret model outputs and maintain appropriate skepticism while remaining open to innovation.
Integration and Future Directions
Building the Continuous Learning Loop
The convergence of AI, nutrition science, and aging research offers the opportunity to link dietary data, biological markers, and clinical outcomes into a continuous learning loop. This vision requires the integration of multiple data streams (including real-time dietary intake, biological age, metabolic responses, and the microbiome) into unified predictive models that constantly improve. Platforms like ZOE exemplify this approach, leveraging machine learning to generate individualized dietary recommendations based on comprehensive biological data. Industry-academia collaborations are also advancing the development of functional foods and smart packaging technologies, demonstrating the practical implementation of AI-informed systems.
Addressing Implementation Challenges
Realizing this vision requires addressing critical challenges through collaboration among clinicians, data scientists, regulators, and patients. For European professionals, the coming years will be crucial to demonstrate that ethical AI can strengthen clinical practice by building trust. Priorities include data standardization to foster interoperability, rigorous clinical validation in diverse populations, ensuring equity to avoid exacerbating health disparities, specific professional training, and patient-centered design that ensures usability and cultural appropriateness.
Research Priorities
Future research should focus on longitudinal validation studies to evaluate long-term outcomes, elucidating the underlying biological mechanisms, comparative effectiveness studies against the standard of care, implementation science to optimize clinical adoption, and economic evaluations considering the cost-effectiveness of AI-enabled care.
Conclusion
Artificial intelligence has evolved from a speculative technology to a practical tool reshaping nutrition and aging research. Deep learning models provide accurate estimates of biological age, apps enable precise dietary assessments, and predictive algorithms identify malnutrition risk early. These advances are current realities supported by solid scientific evidence.
However, realizing the full potential of AI requires vigilance. The European AI Act and the EFSA roadmap establish rigorous standards for oversight and transparency. Nutrition professionals must develop AI literacy as a core competency, learning to critically evaluate tools and keeping human judgment at the center of care. Engaging in this transformation is essential to deliver evidence-based care that extends not only lifespan but, above all, the years lived in good health.
#NutriAI #NutriAINewsletter #ArtificialIntelligence #AI #Nutrition #ScientificCommunication #FoodTech #FoodSafety #AIRegulation #EFSA #RegulatoryCompliance #ISO42001 #HealthClaims #DigitalInnovation #ResponsibleAI #AITransparency #Governance #DataScience #FoodCompliance #DigitalNutrition #FoodLaw #HighRiskAI #TrustInAI #AINews #ScientificCommunication #EUAIAct #MedicalEducation #AIliteracy #ContinuingEducation #Dietitians #Nutritionists #LargeLanguageModels #AIAct #ClinicalDecisionSupport #DigitalHealth #Nutrition
Disclaimer: All rights to images and content used belong to their respective owners. This article is provided for educational and informational purposes only. It does not constitute legal or regulatory advice. Organizations should consult qualified legal and regulatory experts before implementing AI systems in the nutrition sector.
--------------------------------------------------------------------------
Bibliographic and Regulatory References
Wilczok D, Zhavoronkov A, et al. Deep learning and generative artificial intelligence in aging research and healthy longevity medicine. Aging (Albany NY). 2025;17(1):251–275. doi: 10.18632/aging.206190
Abadir PM. The promise of AI and technology to improve quality of life and care for older adults. Nature Aging. 2023;3:239–240. doi: 10.1038/s43587-023-00371-x
Kalargiros G, et al. Towards AI-driven longevity research: an overview. Frontiers in Aging. 2023;4:1057204. doi: 10.3389/fragi.2023.1057204
Lyu YX, Zhavoronkov A, Moskalev A, et al. Longevity biotechnology: bridging AI, biomarkers, geroscience and clinical applications for healthy longevity. Aging (Albany NY). 2024;16(20):13382–13441. doi: 10.18632/aging.206135
Horvath S. DNA methylation age of human tissues and cell types. Genome Biology. 2013;14(10):R115. doi: 10.1186/gb-2013-14-10-r115
Hannum G, et al. Genome-wide methylation profiles reveal quantitative views of human aging rates. Molecular Cell. 2013;49(2):359–367. doi: 10.1016/j.molcel.2012.10.016
Levine ME, et al. An epigenetic biomarker of aging for lifespan and healthspan. Aging (Albany NY). 2018;10(4):573–591. doi: 10.18632/aging.101414
Liu Z, et al. Underlying features of epigenetic aging clocks in vivo and in vitro. Aging Cell. 2020;19(1):e13080. doi: 10.1111/acel.13080
Higgins-Chen AT, et al. A computational solution for bolstering reliability of epigenetic clocks: implications for clinical trials and longitudinal tracking. Nature Aging. 2022;2:644–661. doi: 10.1038/s43587-022-00248-2
Qiu W, et al. ExplaiNAble BioLogical Age (ENABL Age): an artificial intelligence framework for interpretable biological age. The Lancet Healthy Longevity. 2023;4(12):e711–e723. doi: 10.1016/S2666-7568(23)00189-7
Zhang Q. An interpretable biological age. The Lancet Healthy Longevity. 2023;4(12):e662–e663. doi: 10.1016/S2666-7568(23)00213-1
Durão C, et al. Artificial Intelligence Applications to Measure Food and Nutrient Intakes: Scoping Review. Journal of Medical Internet Research. 2024;26:e54557. doi: 10.2196/54557
Agrawal K, et al. Artificial intelligence in personalized nutrition and food manufacturing: a comprehensive review. Frontiers in Nutrition. 2025;12:1636980. doi: 10.3389/fnut.2025.1636980
Mezgebo LB, et al. Accuracy of digital image-based dietary assessment methods compared to human estimation: systematic review. Public Health Nutrition. 2023;26(12):2836–2849. doi: 10.1017/S1368980023001398
Wang X, et al. Artificial Intelligence Applications to Personalized Dietary Recommendations: A Systematic Review. Healthcare. 2025;13(12):1417. doi: 10.3390/healthcare13121417
Sun Y, et al. Artificial intelligence-integrated continuous glucose monitoring for prediabetes management: A comprehensive review. Frontiers in Endocrinology. 2024;15:1320880. doi: 10.3389/fendo.2024.1320880
Contreras I, Vehi J. Artificial intelligence for diabetes management and decision support: literature review. Journal of Medical Internet Research. 2018;20(5):e10775. doi: 10.2196/10775
Zeevi D, et al. Personalized nutrition by prediction of glycemic responses. Cell. 2015;163(5):1079–1094. doi: 10.1016/j.cell.2015.11.001
Wang X, et al. Artificial Intelligence Applications to Personalized Dietary Interventions: Systematic Review. Healthcare. 2025;13(12):1417. doi: 10.3390/healthcare13121417
Deng Z, et al. AI-driven personalized dietary recommendations in type 2 diabetes: A systematic review of machine learning approaches. Diabetes Care. 2024 [in press]. doi: 10.2337/dc23-2274
Güney-Coşkun M, et al. The Future of Artificial Intelligence-driven Personalized Nutrition in Gastroenterology and Hepatology. Journal of Translational Gastroenterology. 2026;5(1). doi: 10.14218/JTG.2025.00043
Hjorth MF, et al. Pre-treatment microbial Prevotella-to-Bacteroides ratio and diet effects on weight loss. International Journal of Obesity. 2019;43:149–157. doi: 10.1038/s41366-018-0093-2
Rinninella E, et al. Artificial Intelligence in Malnutrition: A Systematic Literature Review. Advances in Nutrition. 2024;15(9):100264. doi: 10.1016/j.advnut.2024.100264
Cederholm T, et al. GLIM criteria for the diagnosis of malnutrition: A consensus report from the global clinical nutrition community. Clinical Nutrition. 2019;38(1):1–9. doi: 10.1016/j.clnu.2018.08.002
Deutz NE, et al. Protein intake and exercise for optimal muscle function with aging: recommendations from the ESPEN Expert Group. Clinical Nutrition. 2014;33(6):929–936. doi: 10.1016/j.clnu.2014.04.007
Sun P, et al. Interpretable machine learning-based predictive model for malnutrition in subacute post-stroke patients: an internal and external validation study. Frontiers in Nutrition. 2026;12:1692020. doi: 10.3389/fnut.2025.1692020
Chen L, et al. LASSO-based feature selection for clinical prediction models: methodology and applications. Journal of Clinical Epidemiology. 2020;123:1–12. doi: 10.1016/j.jclinepi.2020.03.015
Sun Q, Che J. Machine Learning-Driven Personalized Risk Prediction: Developing an Explainable Sarcopenia Model for Older European Adults with Arthritis. Journal of Clinical Medicine. 2026;15(3):892. doi: 10.3390/jcm15030892
Cruz-Jentoft AJ, et al. Sarcopenia: revised European consensus on definition and diagnosis. Age and Ageing. 2019;48(1):16–31. doi: 10.1093/ageing/afy169
Ren SS, et al. Machine Learning-Based Prediction of In-Hospital Complications in Elderly Patients Using GLIM-, SGA-, and ESPEN 2015-Diagnosed Malnutrition as a Factor. Nutrients. 2022;14(15):3035. doi: 10.3390/nu14153035
Cederholm T, et al. ESPEN guidelines on definitions and terminology of clinical nutrition. Clinical Nutrition. 2017;36(1):49–64. doi: 10.1016/j.clnu.2016.09.004
Kondrup J, et al. Nutritional risk screening (NRS 2002): a new method based on an analysis of controlled clinical trials. Clinical Nutrition. 2003;22(3):321–336. doi: 10.1016/S0261-5614(02)00214-5
European Parliament and Council. Regulation (EU) 2024/1689 of 13 June 2024 laying down harmonised rules on artificial intelligence (Artificial Intelligence Act). Official Journal of the European Union. 2024;L 1689. https://eur-lex.europa.eu/legal-content/EN/TXT/?uri=OJ:L_2024_1689
Gesund.ai. The EU AI Act is Redefining High-Risk Healthcare AI. Industry Regulatory Report. November 2024. https://gesund.ai/blog/eu-ai-act_6KenBl2z3htfqYnAnyIdhn
Ozturk A, et al. The AI Act: responsibilities and obligations for healthcare professionals and organizations. Diagnostic and Interventional Radiology. 2025;31(3). doi: 10.4274/dir.2024-24712
Bersani G, et al. Roadmap for actions on artificial intelligence for evidence management in risk assessment. EFSA Supporting Publications. 2022;19(5):EN-7339. doi: 10.2903/sp.efsa.2022.EN-7339
European Food Safety Authority (EFSA). Roadmap for actions on artificial intelligence for evidence management in risk assessment. May 2022. https://www.efsa.europa.eu/en/supporting/pub/en-7339
EFSA Scientific Committee. Guidance on communication of uncertainty in scientific assessments. EFSA Journal. 2019;17(1):5520. doi: 10.2903/j.efsa.2019.5520
Dahl WJ, et al. Precision nutrition: hype or hope? Advances in Nutrition. 2023;14(6):1349–1363. doi: 10.1016/j.advnut.2023.09.001
Contact details
Follow me on LinkedIn
Nutri-AI 2025 - Alessandro Drago. All rights reserved.
e-mail: info@nutri-ai.net